Methodology
This section provides technical details regarding the algorithm used for the
detection of 1D anomalies.
Anomalies are identified from the detection of maximum, minimum and inflection
points calculated from the first and second order derivatives of individual
data channels. The algorithm relies on the
Numpy.fft
routine for the calculation of derivatives in the Fourier domain.
Detection parameters are available for filtering and grouping co-located
anomalies. The selection process is done in the following order:
Primary detection
Loop over the selected data channels:
Apply the Minimum Data Value threshold.
For every maximum (peak) found on a profile, look on either side for
inflection and minimum points. This forms an anomaly.
Keep all anomalies larger than the Minimum Amplitude
Grouping
Anomalies found along individual data channels are grouped based on spatial
proximity:
Find all peaks within the Maximum Peak Migration distance. The nearest peak is
used if multiple peaks are found on a single channel.
Create an anomaly group that satisfies the following criteria:
Minimum Amplitude
property PeakFinderParams. min_amplitude : int
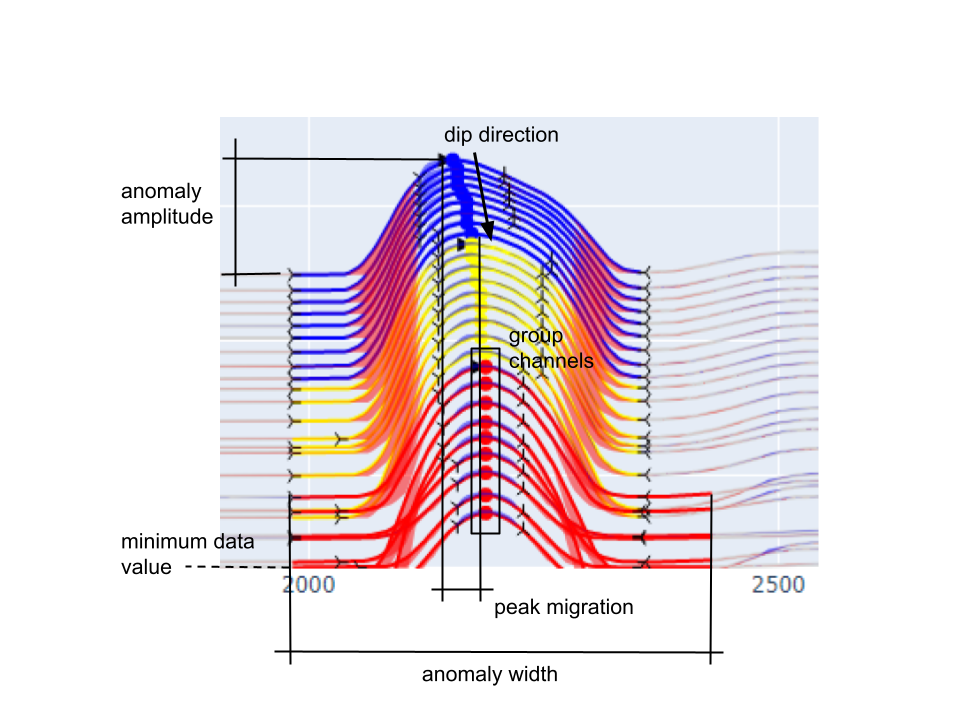
Threshold on the minimum amplitude of the anomaly, expressed as
a percent of the height scaled by the minimum value.
Threshold value (\(\delta A\) ) for filtering small anomalies based on the anomaly
minimum (\(d_{min}\) ) and maximum (\(d_{max}\) ).
\[\delta A = \left|\left|\frac{d_{max} - d_{min}}{d_{min}}\right|\right| \cdot 100\]
See figure
Todo
Add figure showing the effect on anomaly identification
Minimum Data Value
property PeakFinderParams. min_value : float
Minimum absolute data value to be considered for anomaly detection.
The minimum data threshold (\(\delta_d\) ) (see Figure
\[\begin{split}\begin{equation}
d_i =
\begin{cases}
d_i & \;\text{for } d_i > \delta_d \\
nan & \;\text{for } d_i \leq \delta_d\\
\end{cases}
\end{equation}\end{split}\]
Todo
Add figure showing the effect on anomaly identification
Minimum Width
property PeakFinderParams. min_width : float
Minimum anomaly width (m) measured between start and end of bounding minima.
Todo
Add figure showing the effect of anomaly identification
See figure
Maximum Peak Migration
property PeakFinderParams. max_migration : float
Threshold on the lateral shift (m) of peaks within a grouping of anomalies.
Todo
Add figure showing the effect of anomaly identification
See figure
Minimum number of channels
property PeakFinderParams. min_channels : int
Minimum number of data channels required to form a group.
Todo
Add figure showing the effect of anomaly identification
See figure
Merge N Peaks
property PeakFinderParams. n_groups : int
Number of consecutive peaks to merge into a single anomaly.
Todo
Add figure showing the effect of anomaly identification
Max Group Separation
property PeakFinderParams. max_separation : float
Maximum separation between peaks to merge into single anomaly.
Todo
Add figure showing the effect of anomaly identification
Smoothing
property PeakFinderParams. smoothing : int
Number of neighbors used in running mean smoothing.
The running mean replaces each data by the average of it’s N
\[d_i = \frac{1}{N}\sum_{j=-\frac{N}{2}}^{\frac{N}{2}}d_{i+j}\]
where averaging becomes one sided at both ends of the profile. The result is a
smoothed data set where the degree of smoothing scales with the number of
neighbours used in the mean.
Todo
Add reference figure shown for plot residuals.
Show residual
Option to show the positive (blue) and negative (red) residual
Masking Data
property PeakFinderParams. masking_data : geoh5py.data.data.Data | None
Mask object to focus peak finding within an area of interest.